TreesFormer: Multimodal Grammar-Based 3D Tree Reconstruction from Sparse Geodata
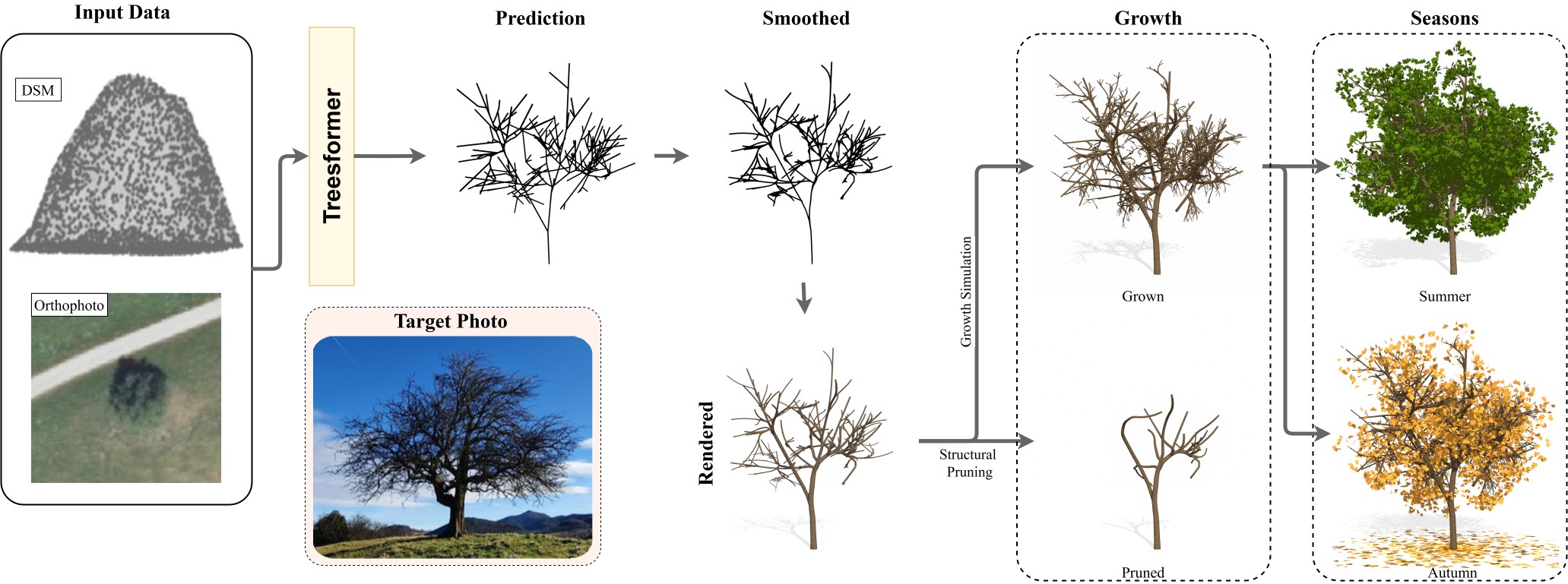

We present TreesFormer, the first grammar-based framework for reconstructing hierarchical 3D tree structures directly from sparse top-down geodata using only a single orthophoto and its corresponding Digital Surface Model (DSM). It employs a multimodal autoregressive transformer that predicts compact parametric L-system grammars from DSM point clouds and orthophoto features, jointly predicting symbolic structure and geometric parameters while enforcing grammar constraints during decoding. To enable supervision in the absence of real-world grammar annotations, we introduce a synthetic multimodal dataset of procedurally generated trees with aligned aerial inputs and ground-truth L-system labels. Experiments show that DSMs drive overall geometric accuracy and crown shape, while orthophoto conditioning improves structural regularity and branching depth; their combination consistently outperforms either modality alone. The model generalizes to real-world Austrian and French aerial data, producing interpretable branching structures suitable for large-scale rural 3D mapping.
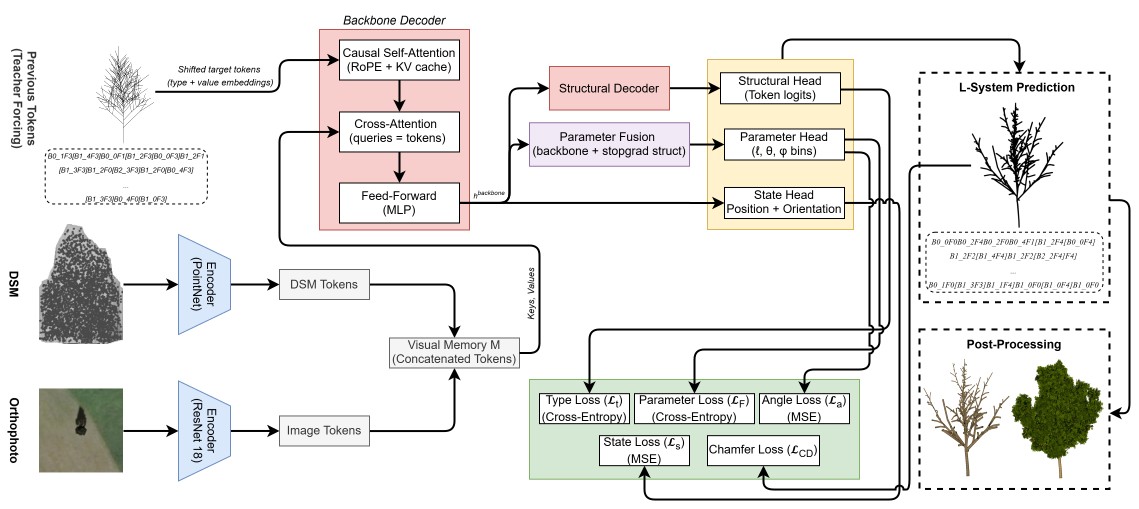
Figure 2: Overall architecture of the multimodal L-system generator. A multimodal visual encoder conditions an autoregressive backbone decoder factorized into a structural branch for token prediction and a parameter branch for geometric prediction, with supervision (green).
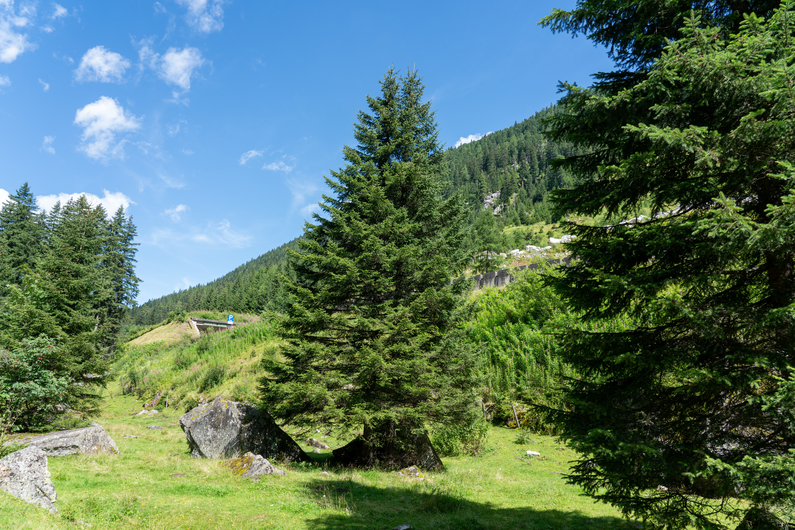
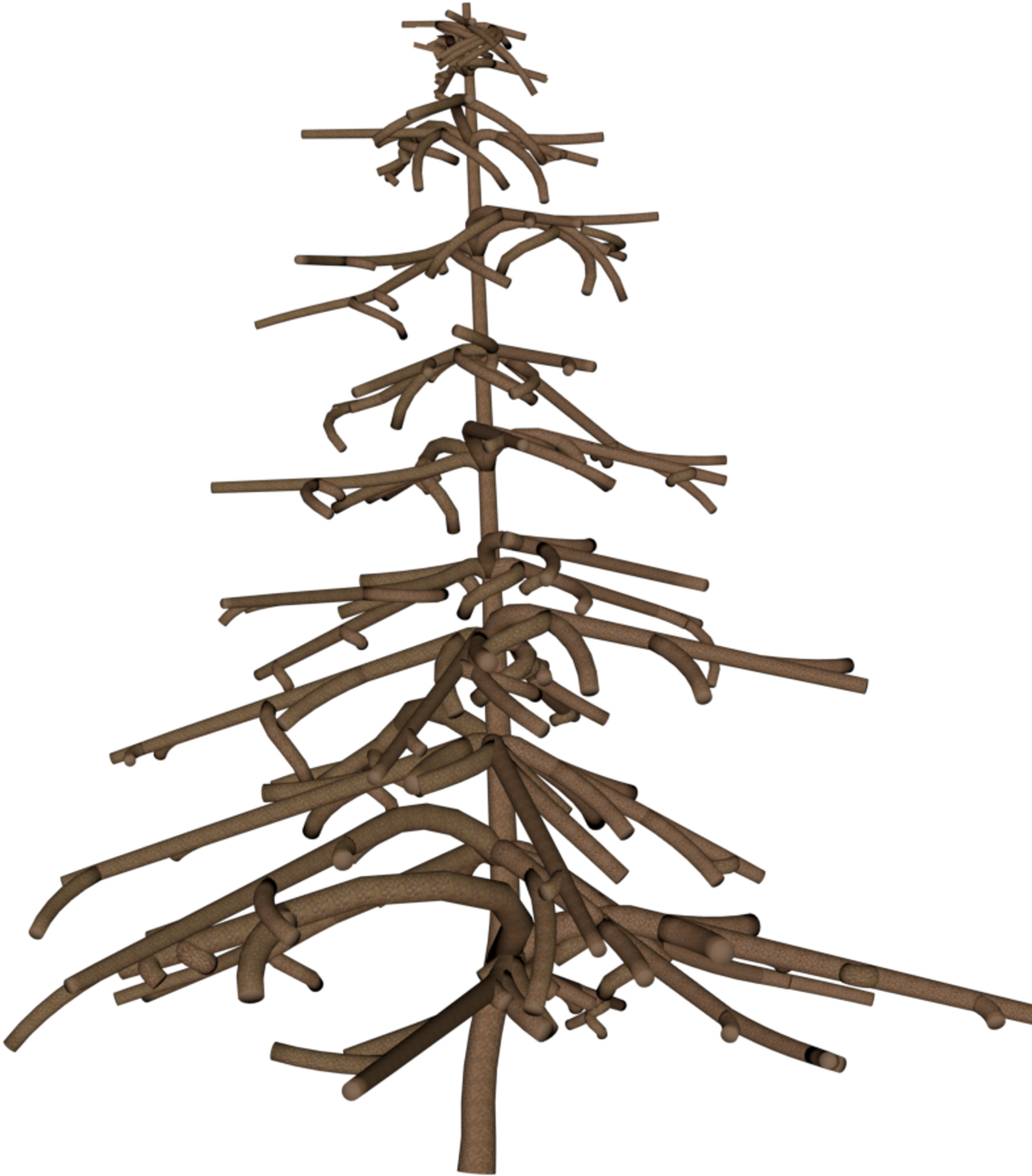
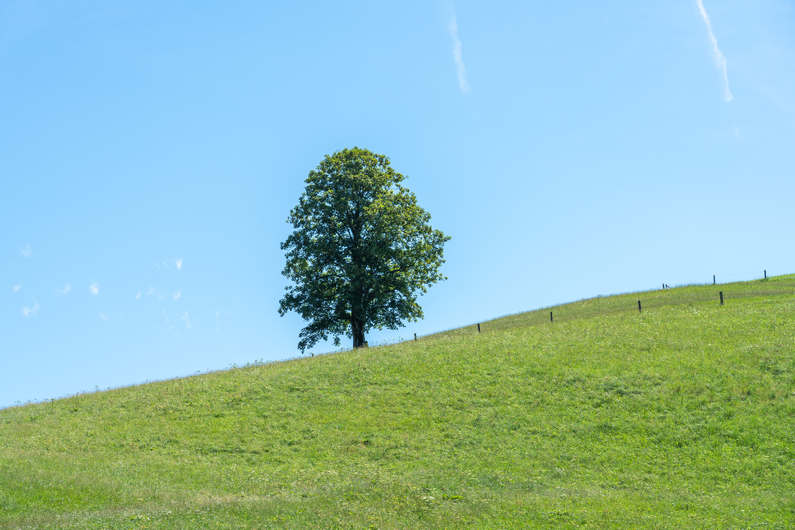
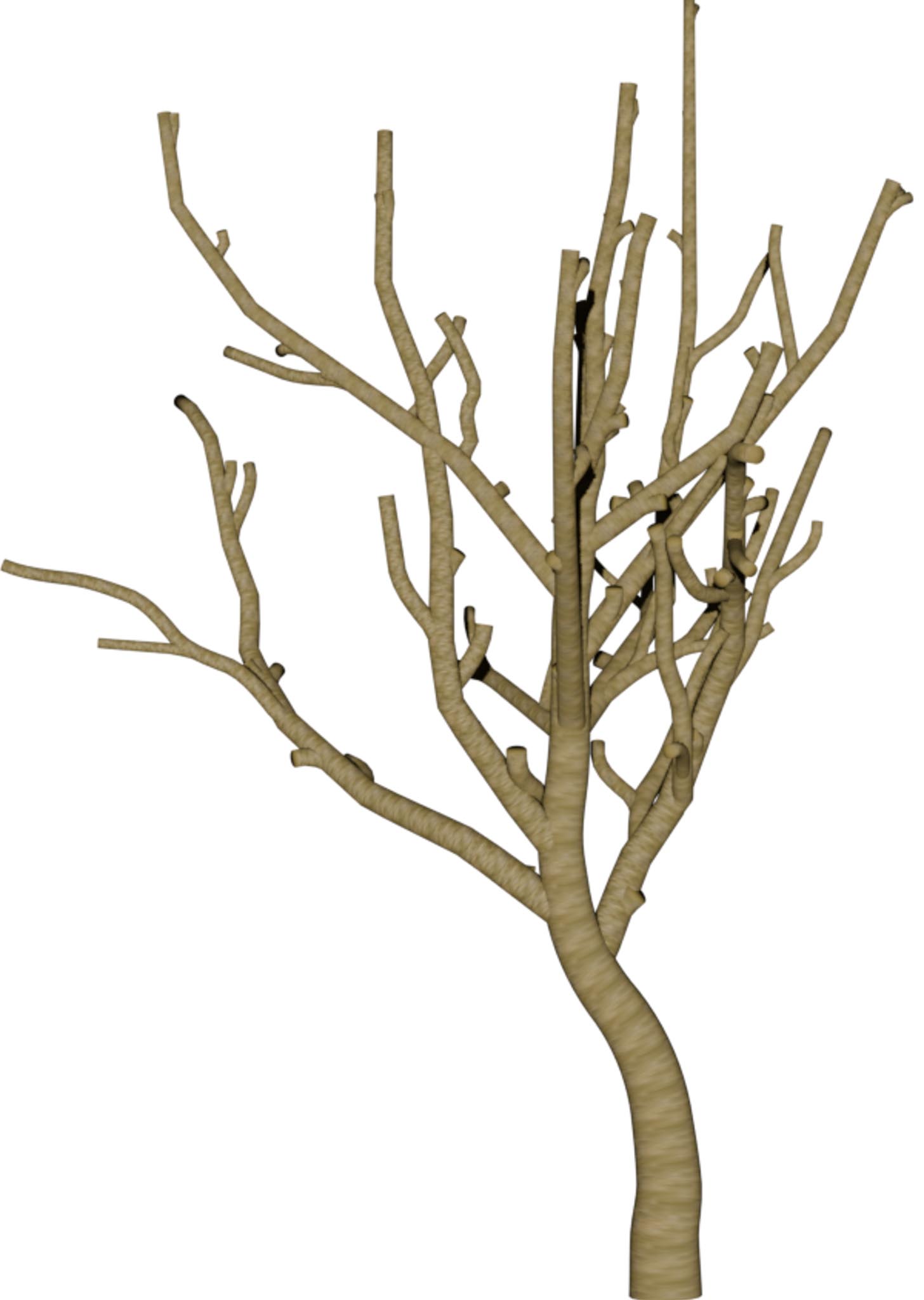
Reconstructions from real-world Austrian landmark trees and French IGN data
| Typical Reconstructions | Difficult Cases | ||||||||
|---|---|---|---|---|---|---|---|---|---|
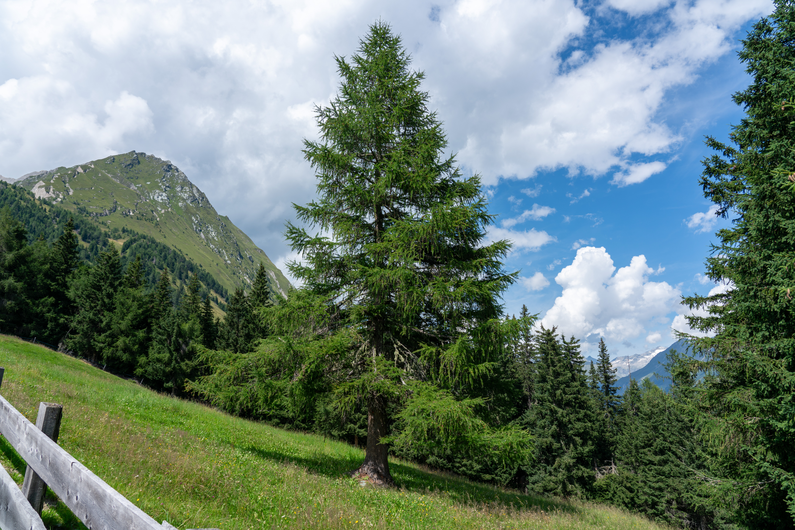
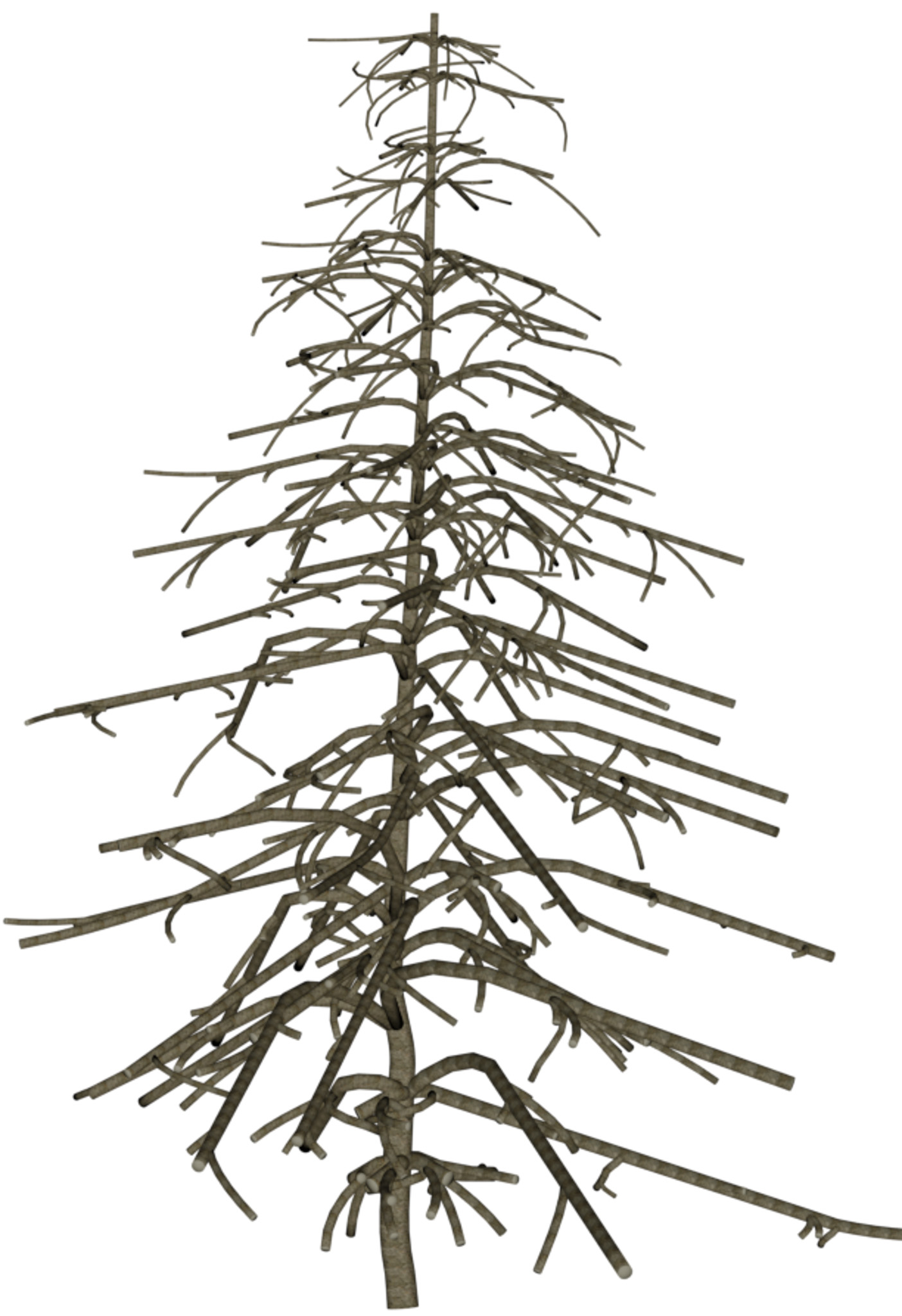
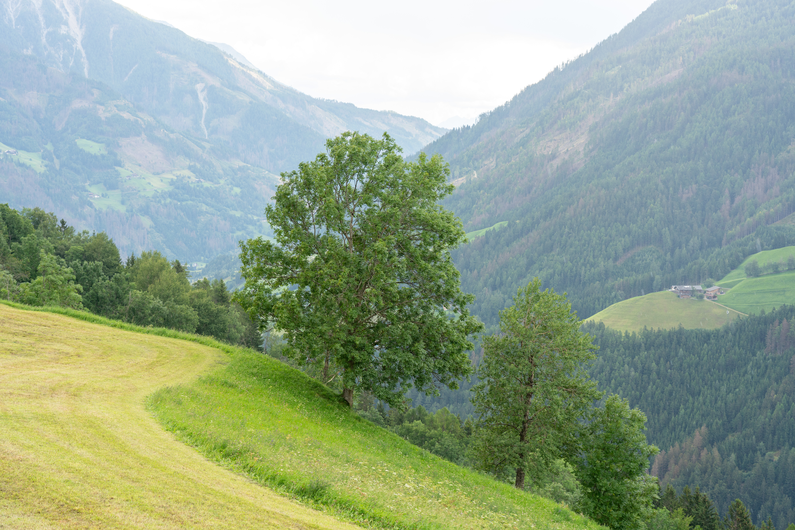
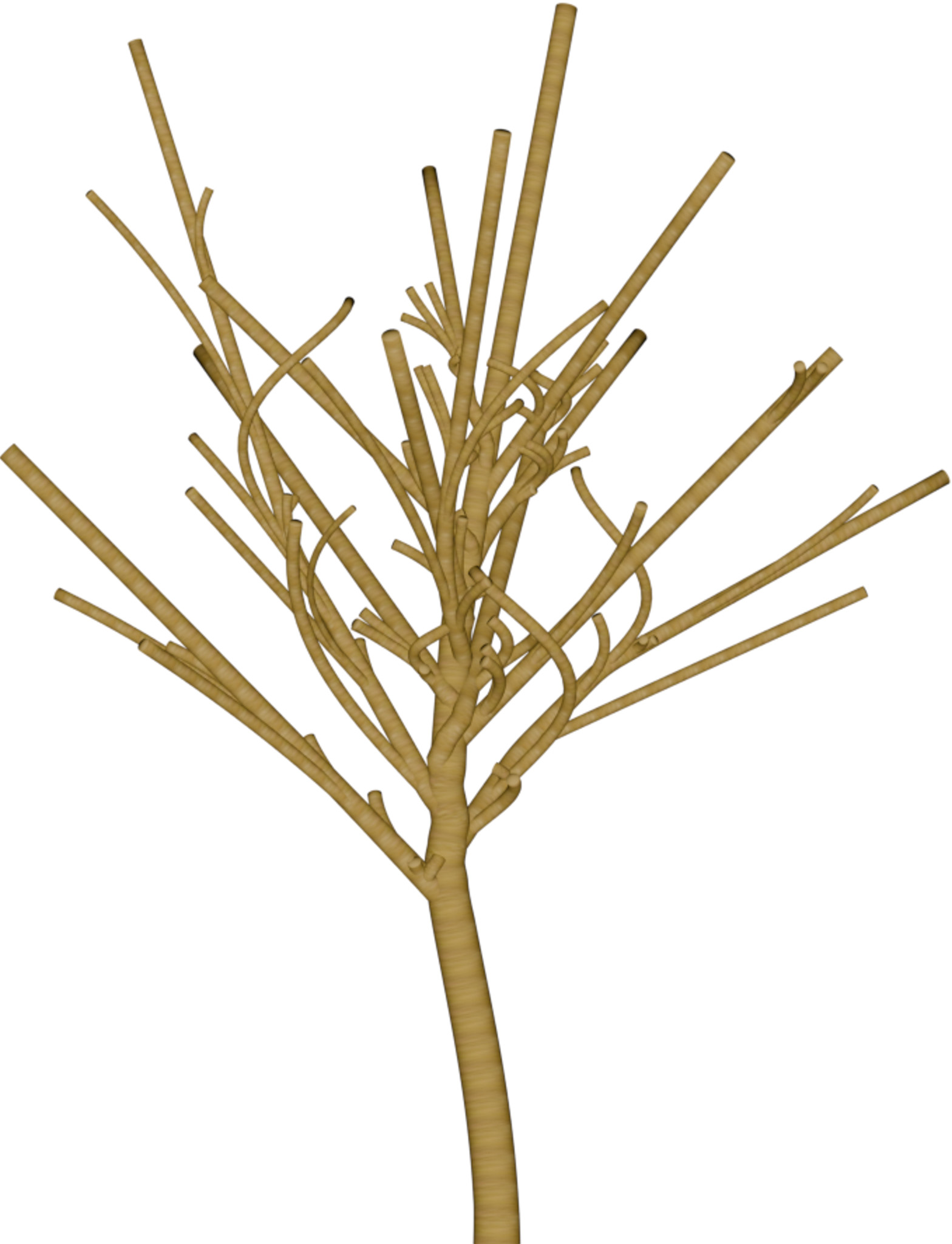
| DSM | Orthophoto | Target | Output | DSM | Orthophoto | Target | Output | ||
| 1 |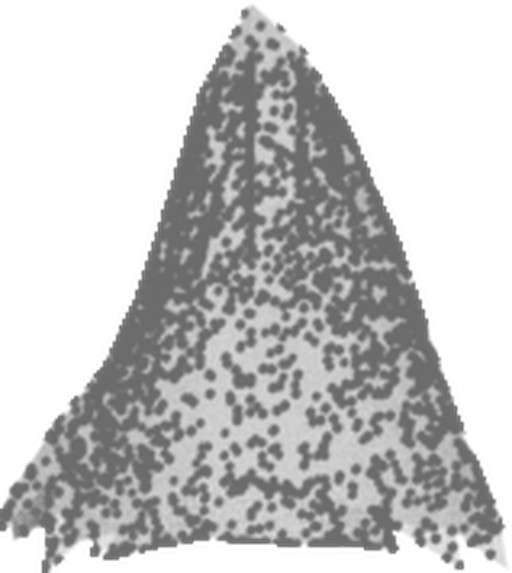  |
 |
 |
 |
1 |  |
 |
 |
 |
| 2 |  |
 |
 |
 |
2 |  |
 |
 |
 |
| 3 |  |
 |
 |
 |
3 |  |
 |
 |
 |
| 4 |  |
 |
 |
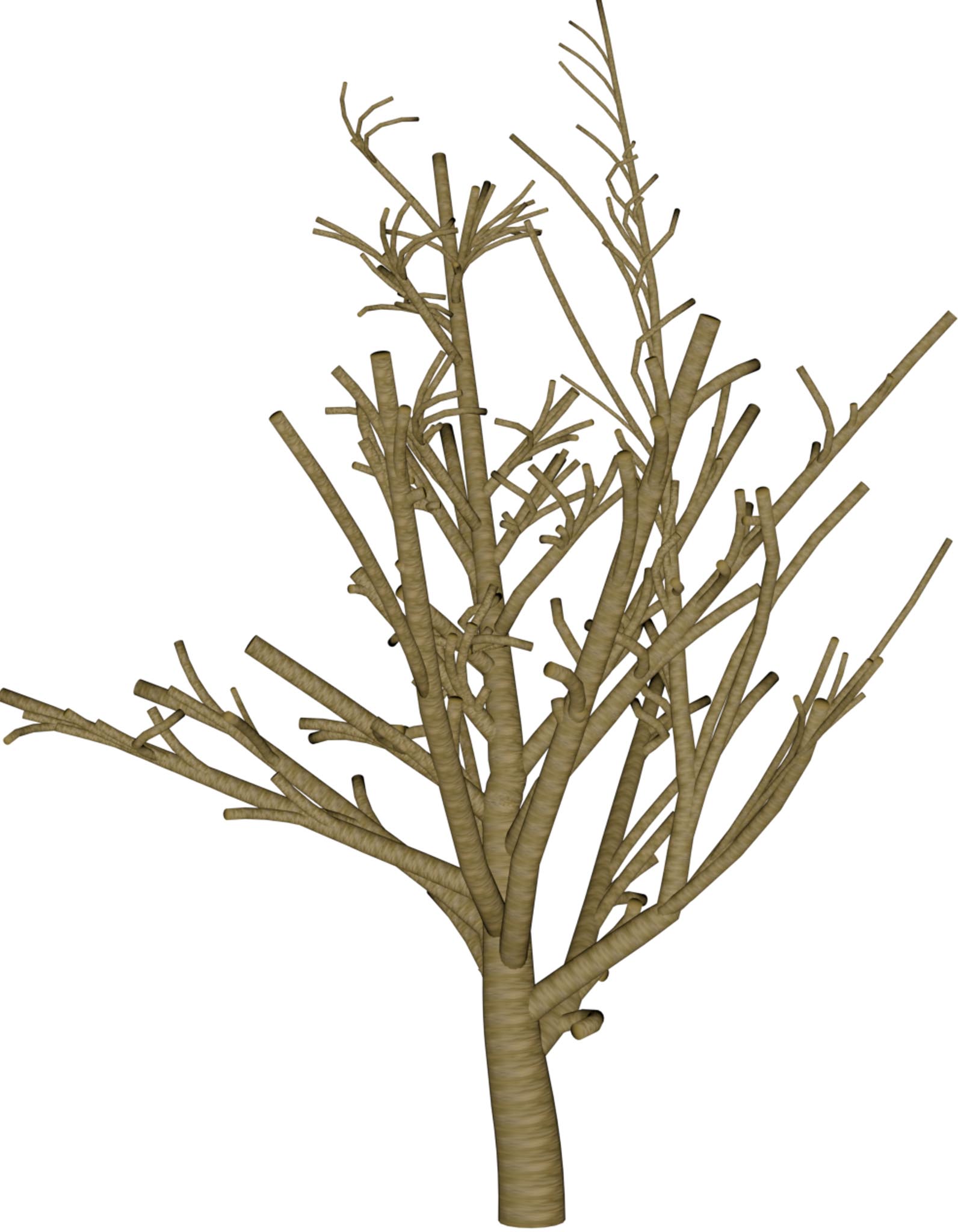 |
4 |  |
 |
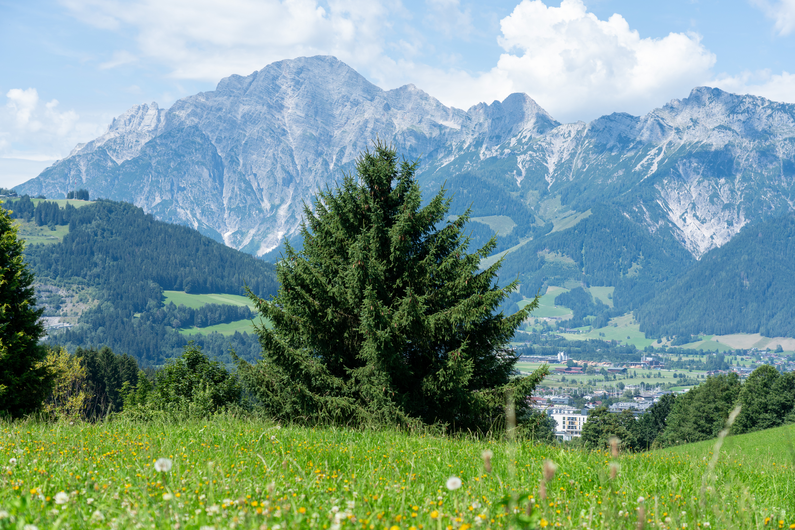 |
 |
Table 1: Reconstructed landmark trees (left) and difficult cases (right). Difficult cases: (1) DSM underestimates tree size, (2) cluttered orthophoto, (3) dead tree, (4) failure -- simultaneous modality ambiguity.
| Inputs | Ground Truth |
Networks | ||||||
|---|---|---|---|---|---|---|---|---|
| DSM | Ortho | Tree D-Fusion |
SVDTree | Latent L-Systems |
TreeON | Ours | ||
| [Lee et al. 2024] | [10656708] | [10.1145/3627101] | [Grammatikaki et al. 2026] | |||||
 |
 |
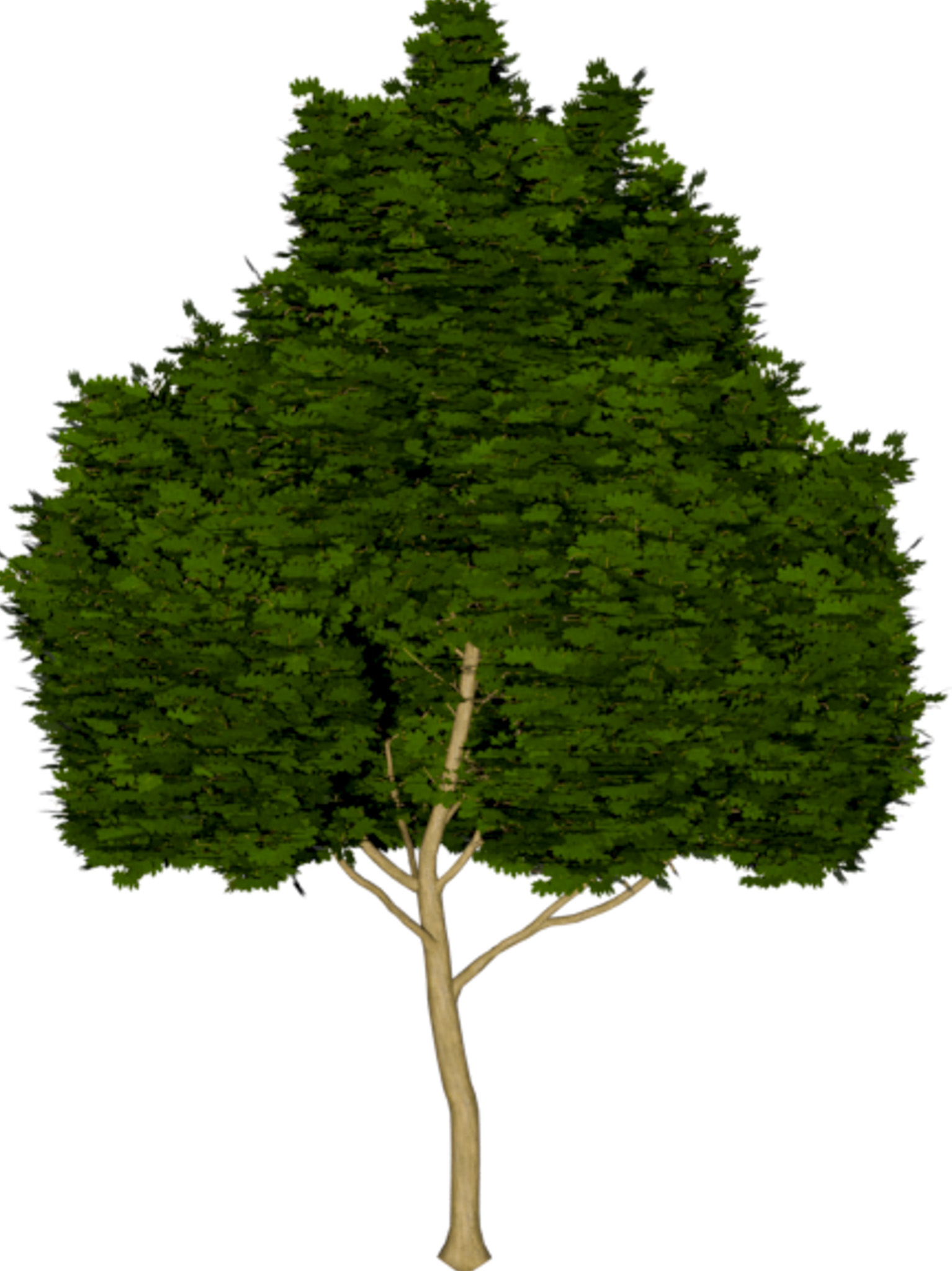 |
 |
 |
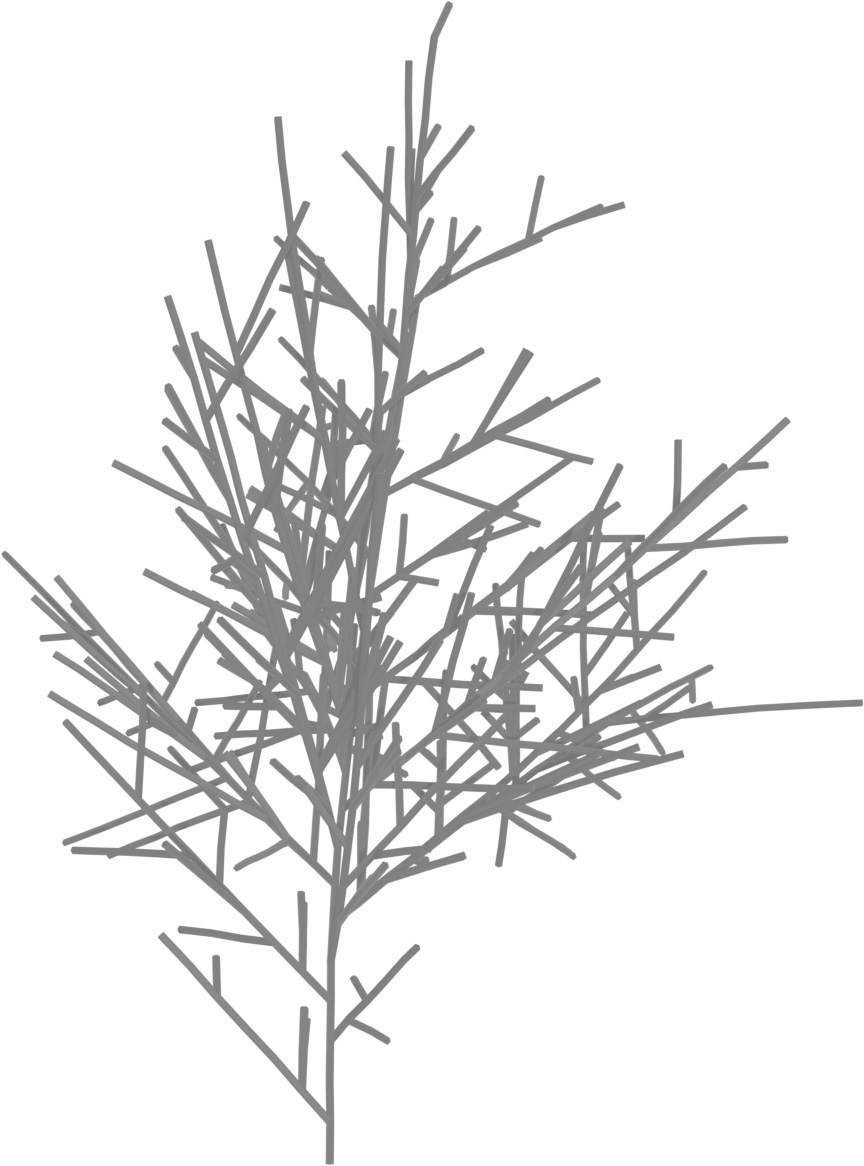 |
 |
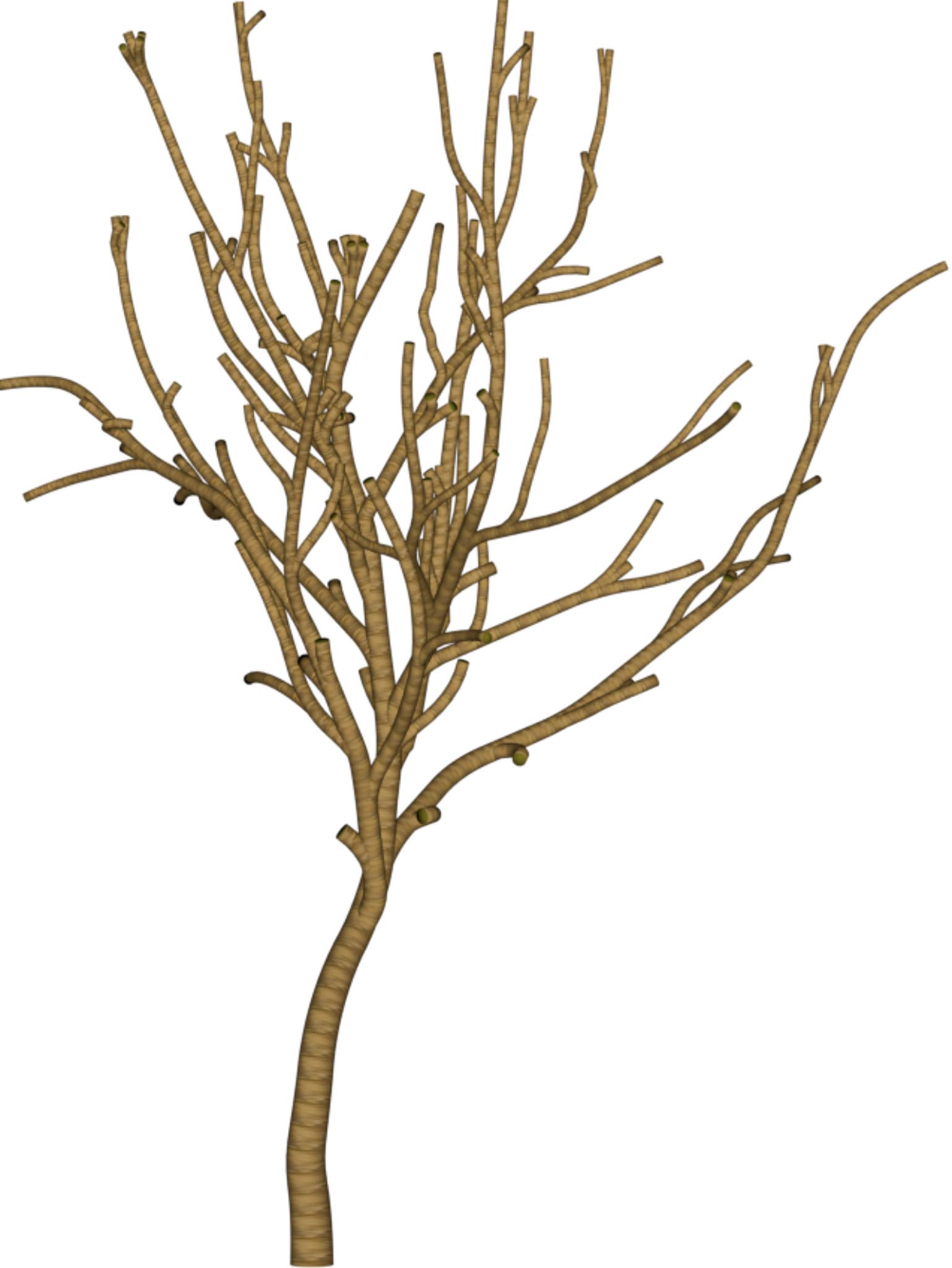 |
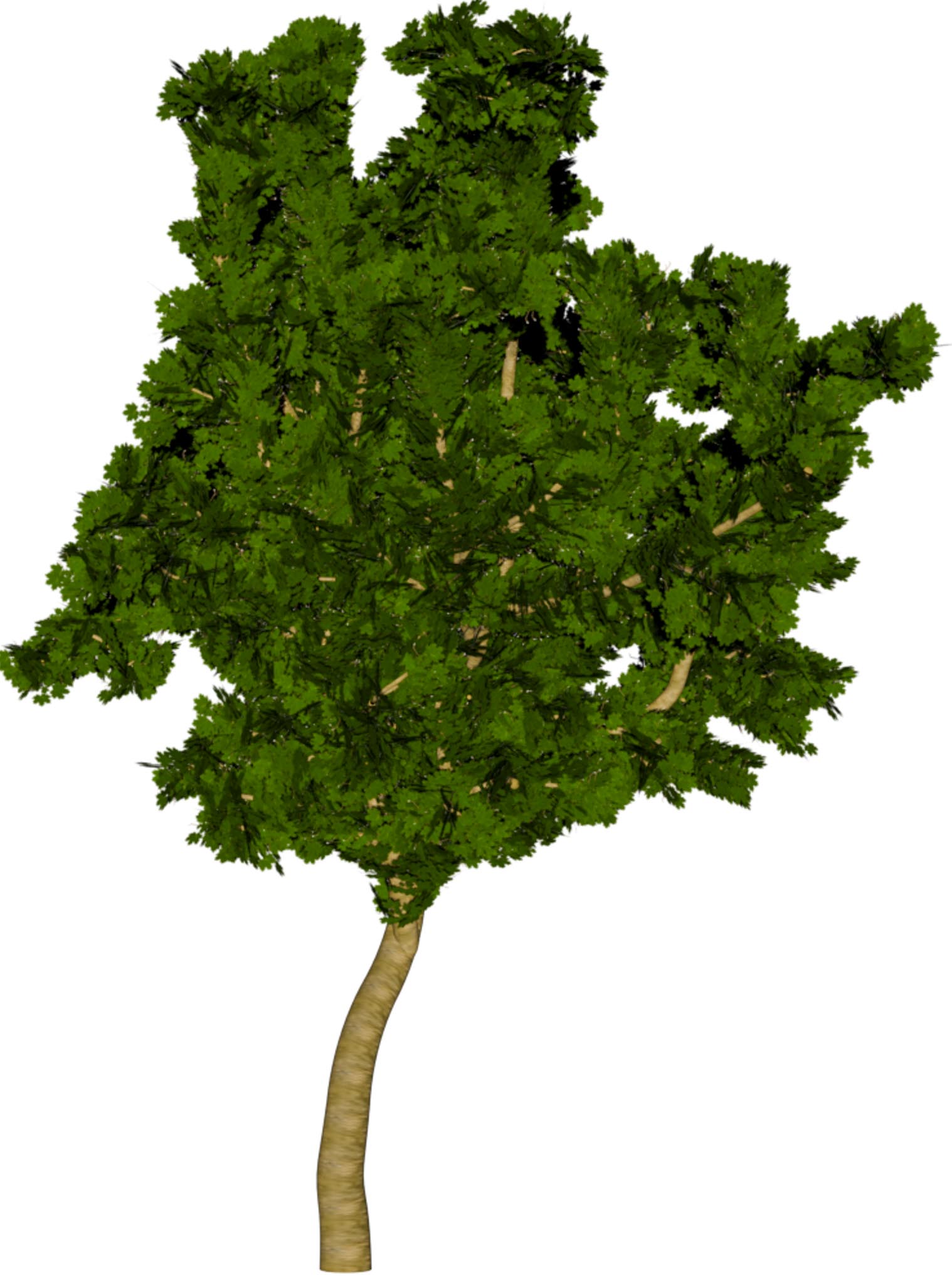 |
 |
 |
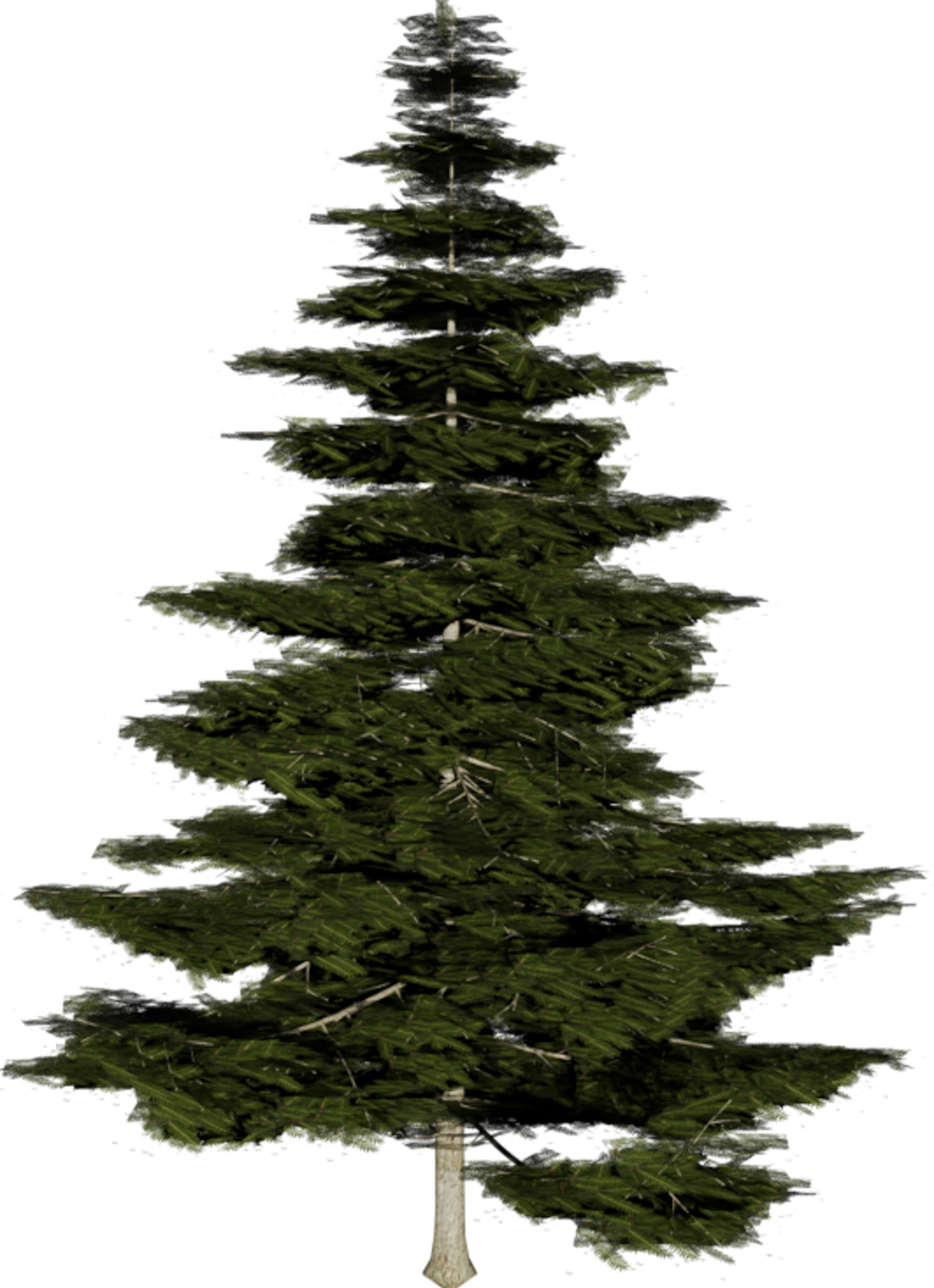 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
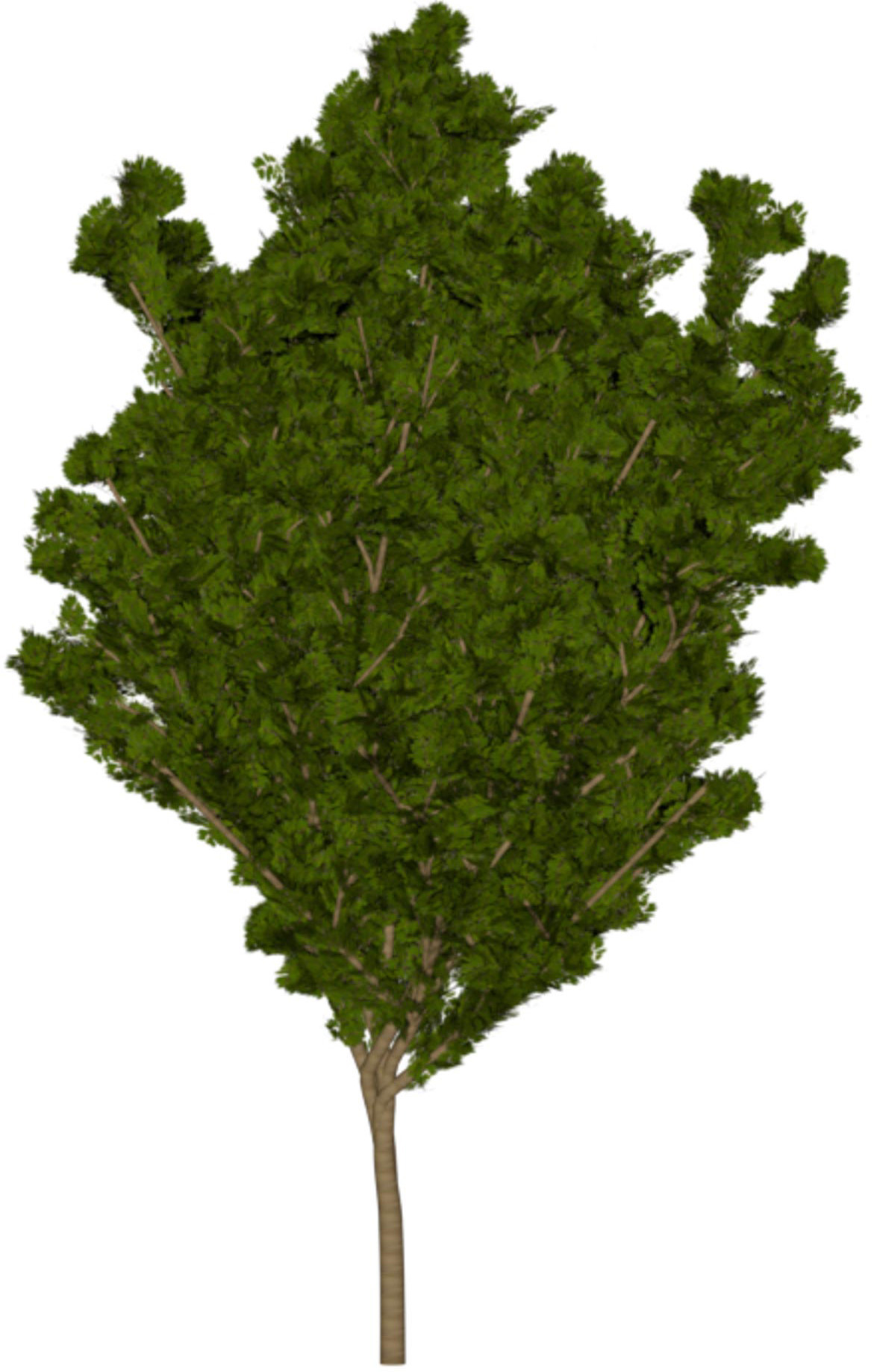 |
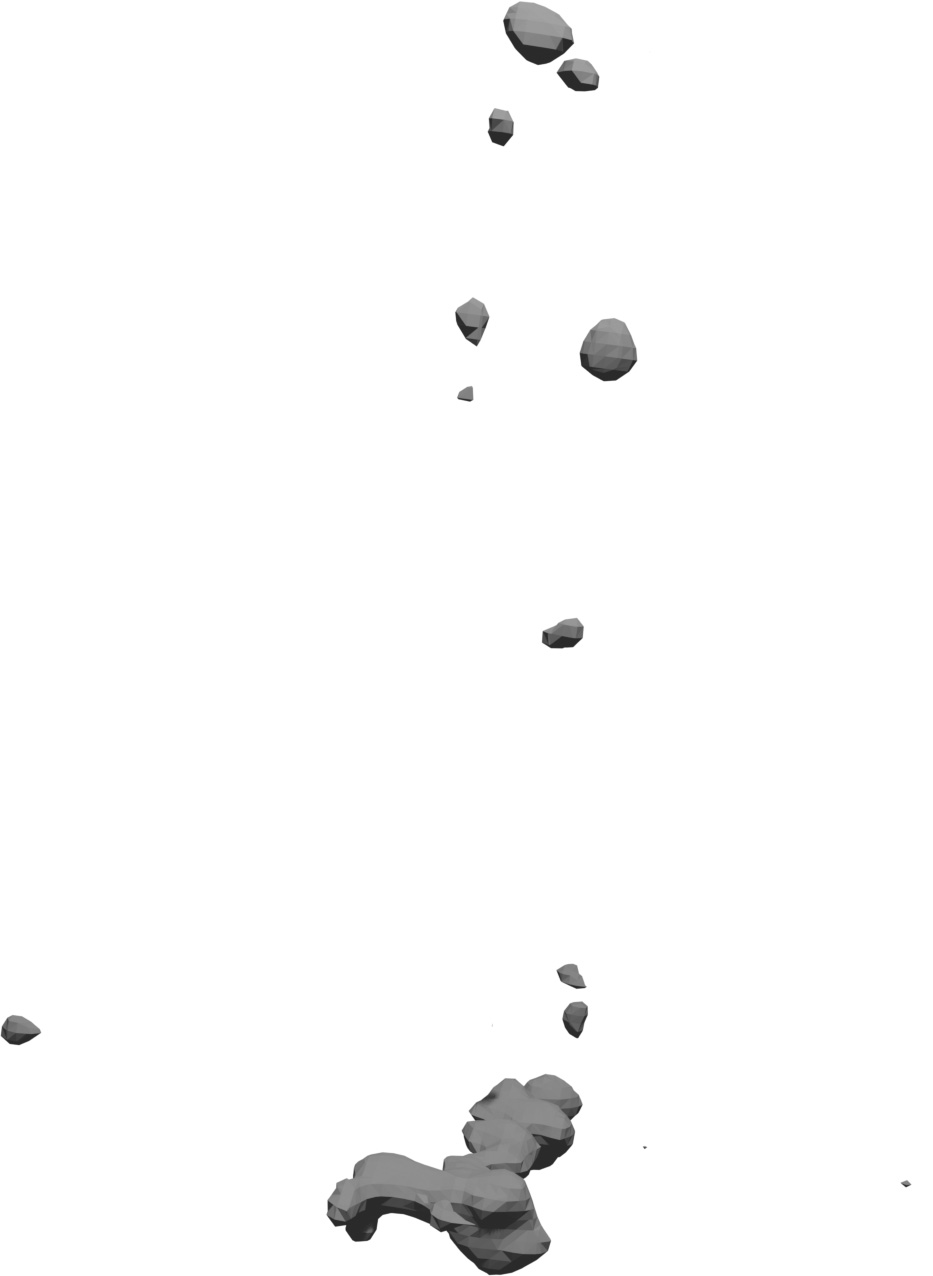
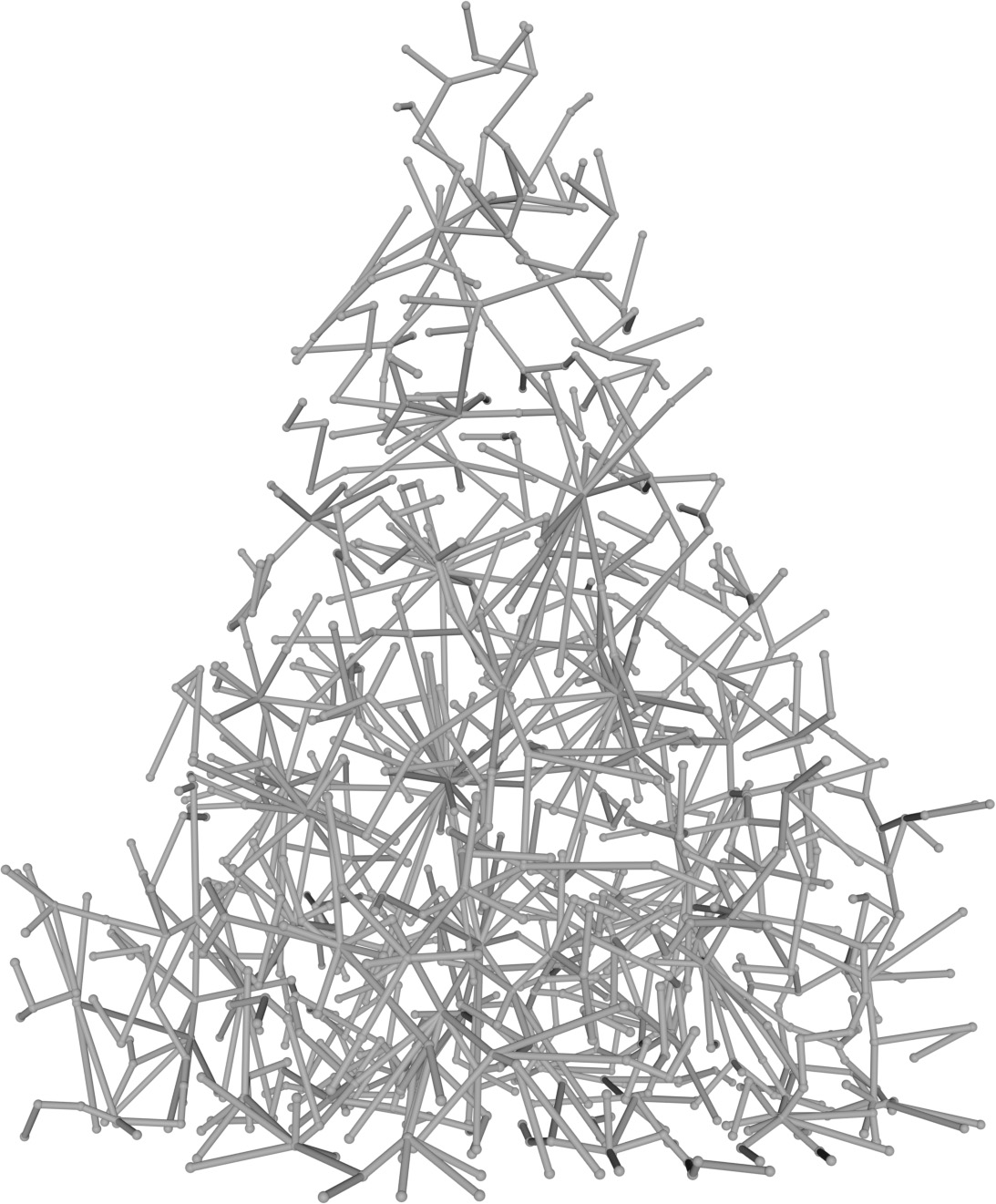
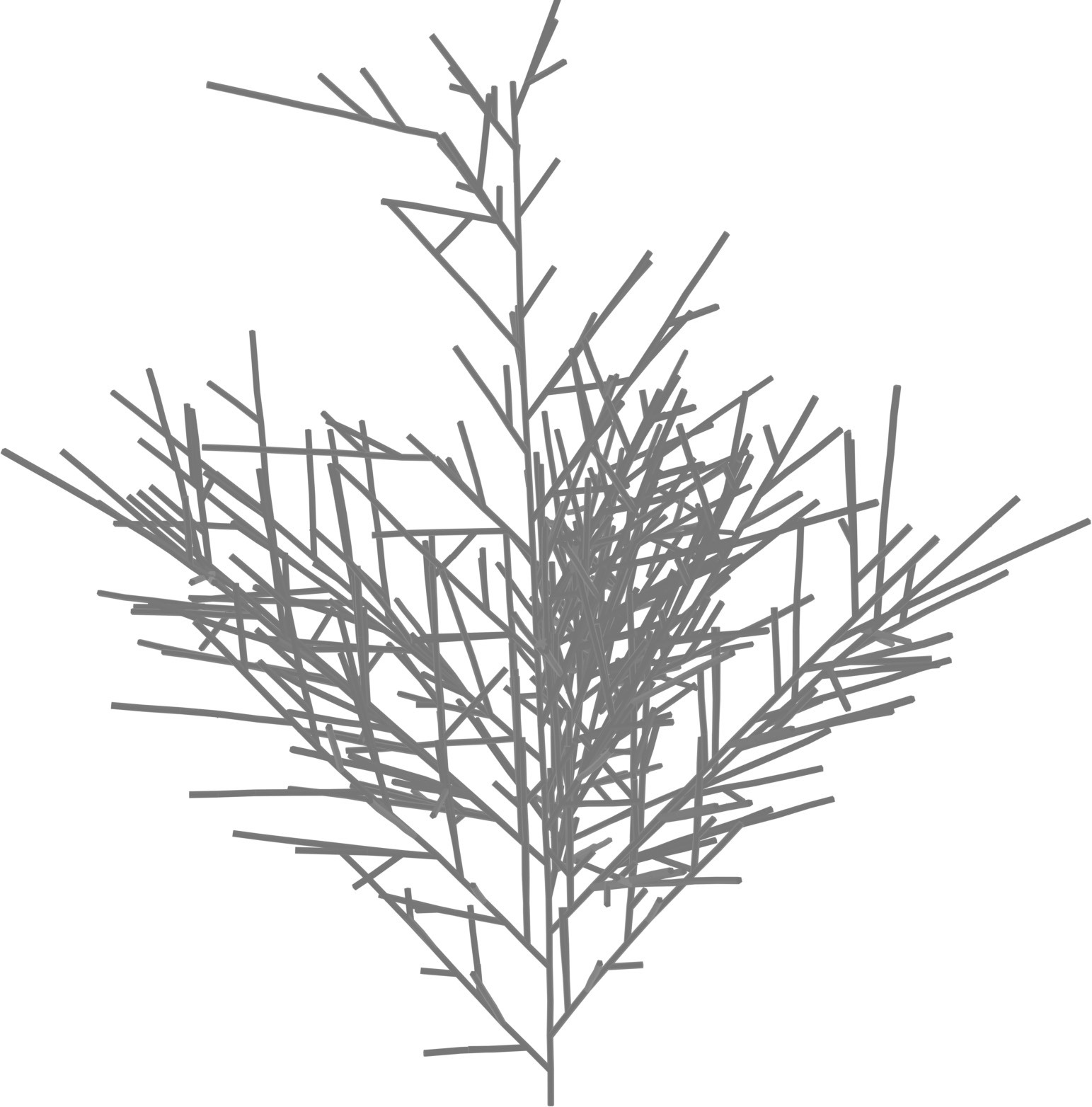
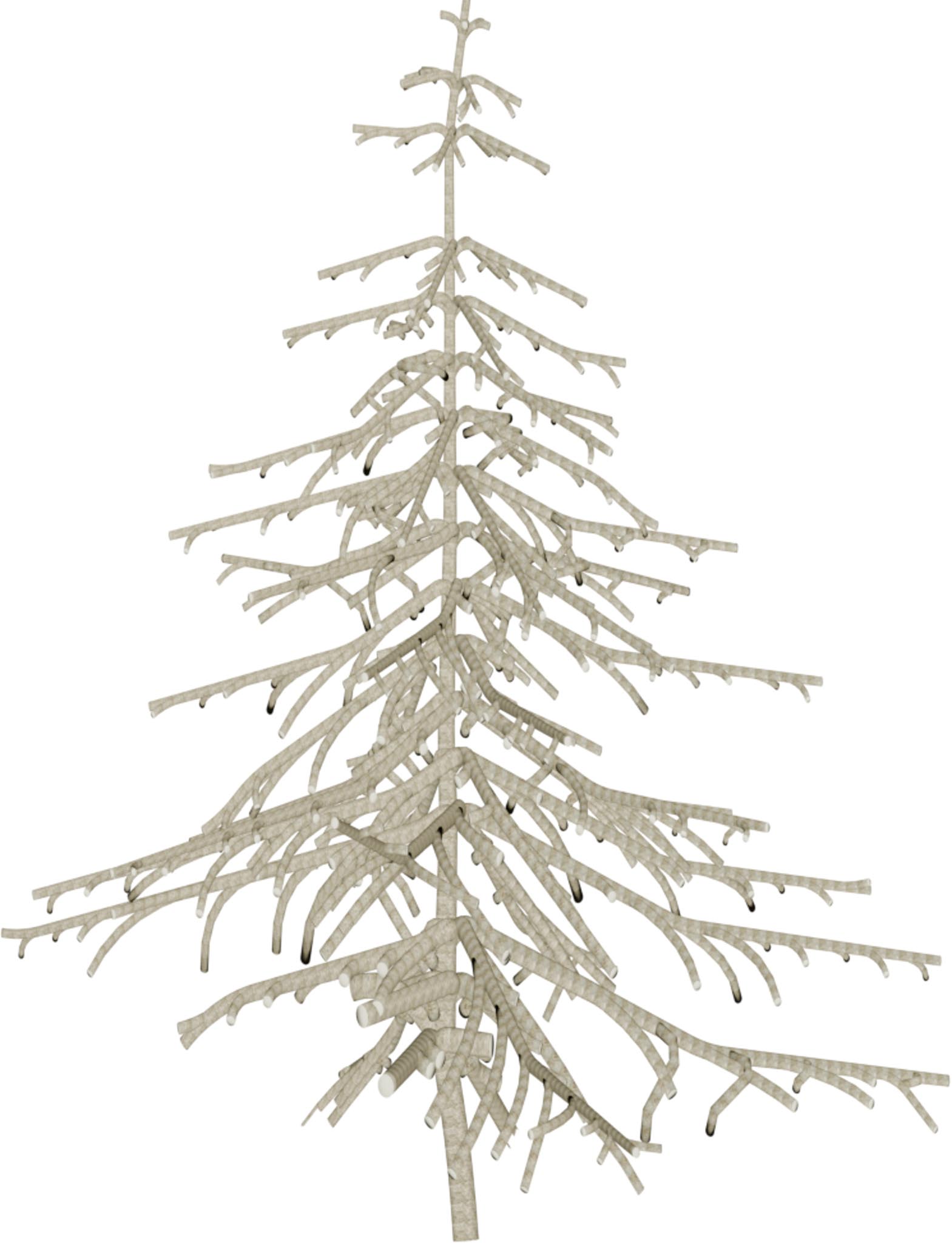
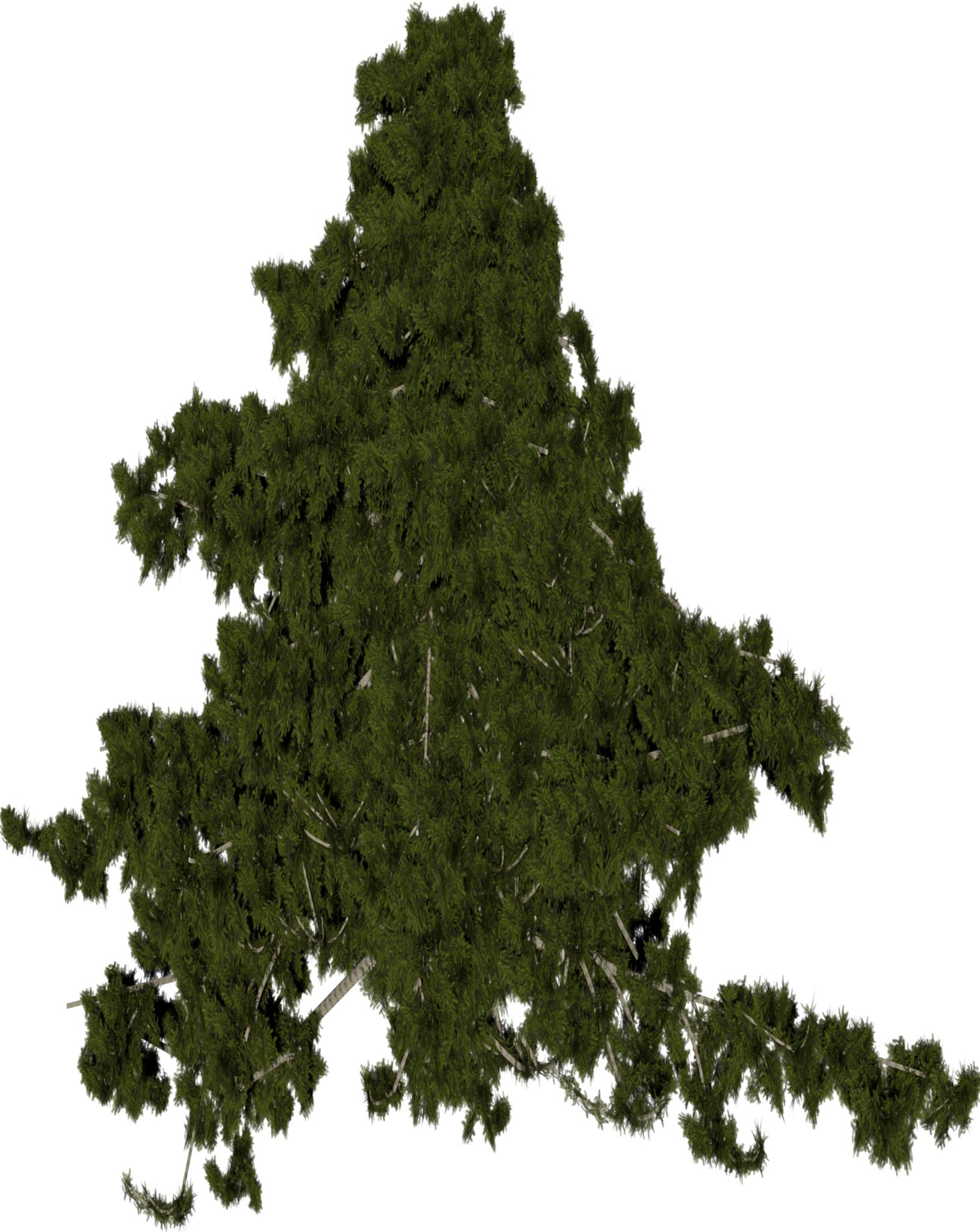
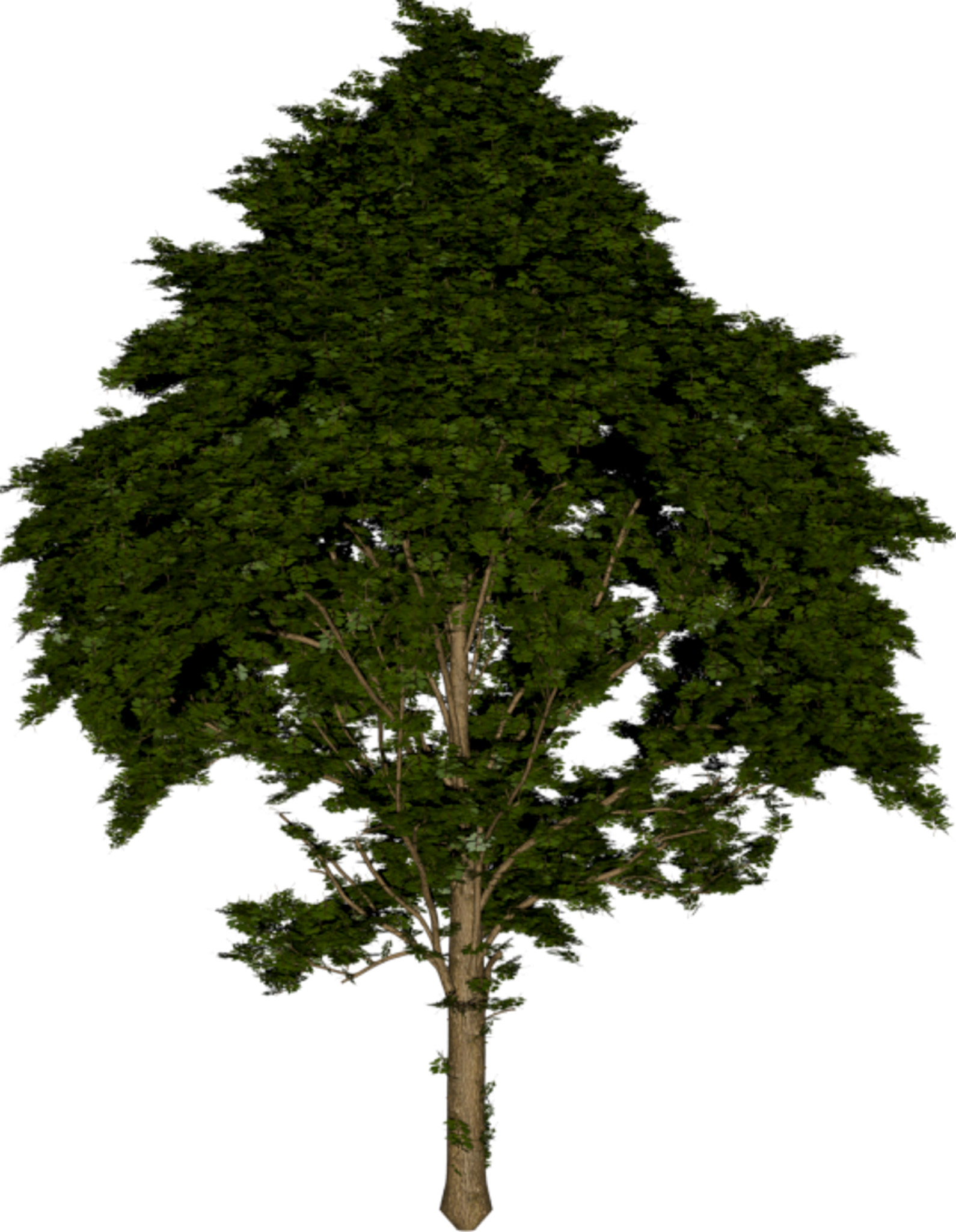
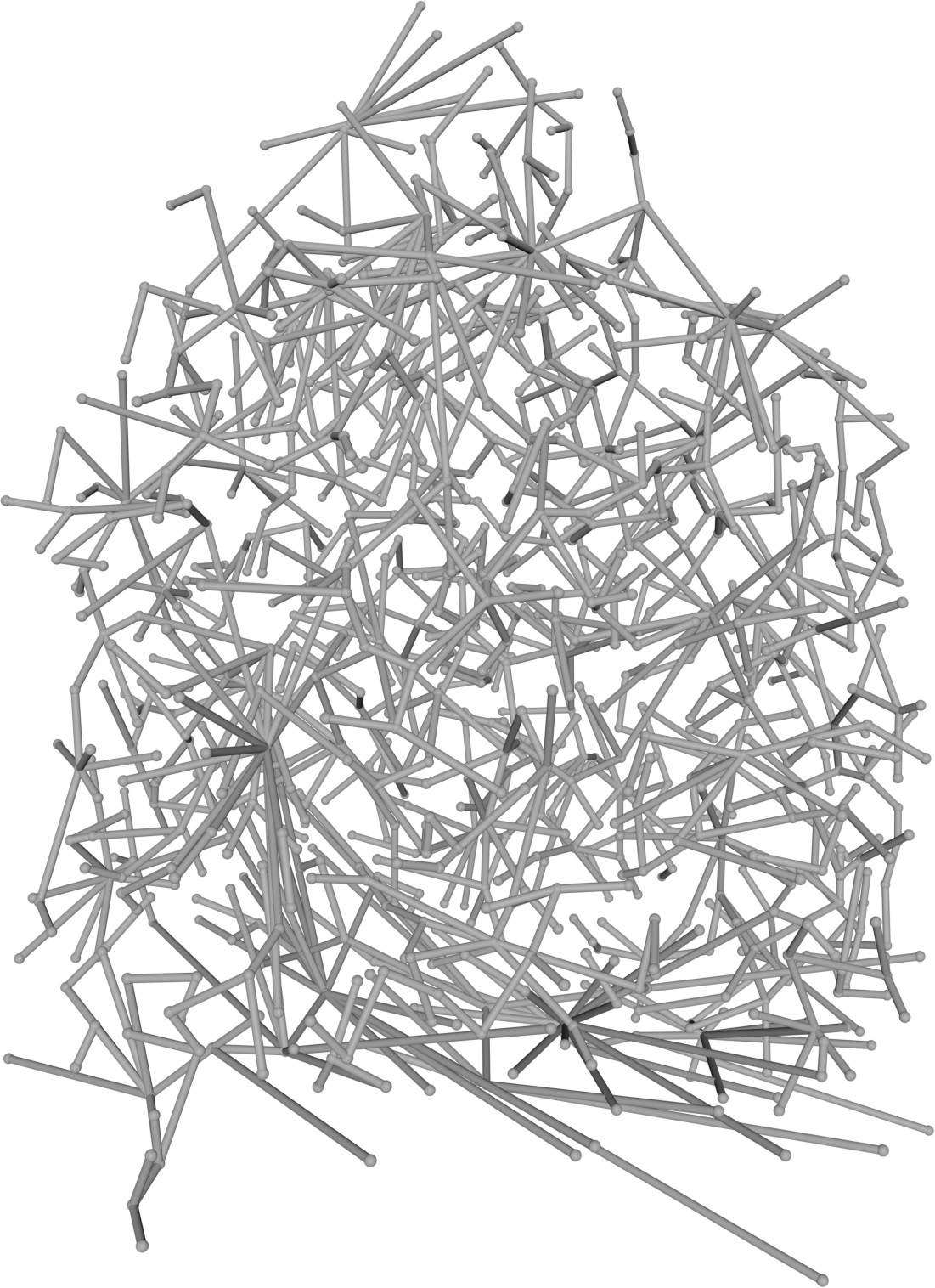
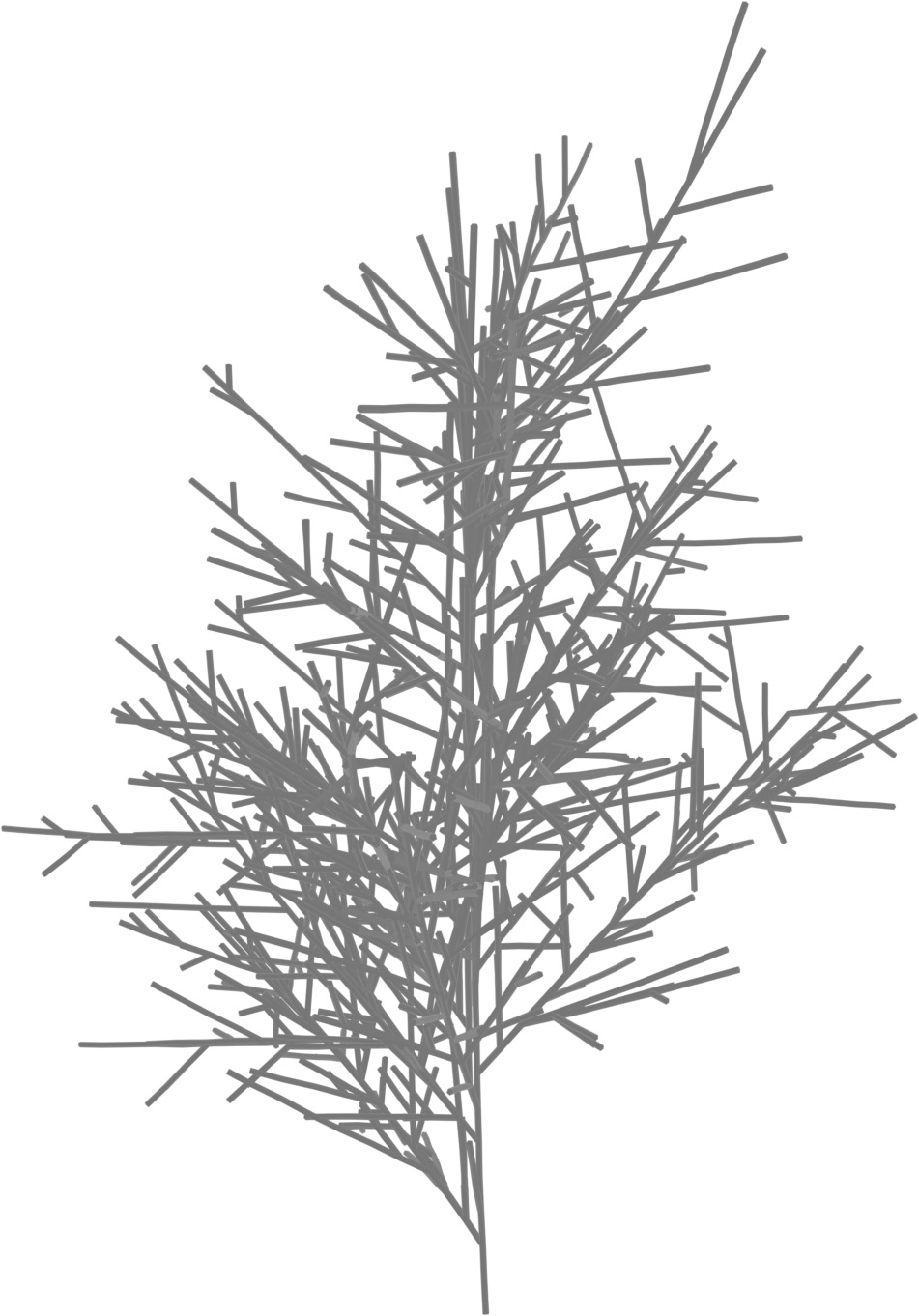
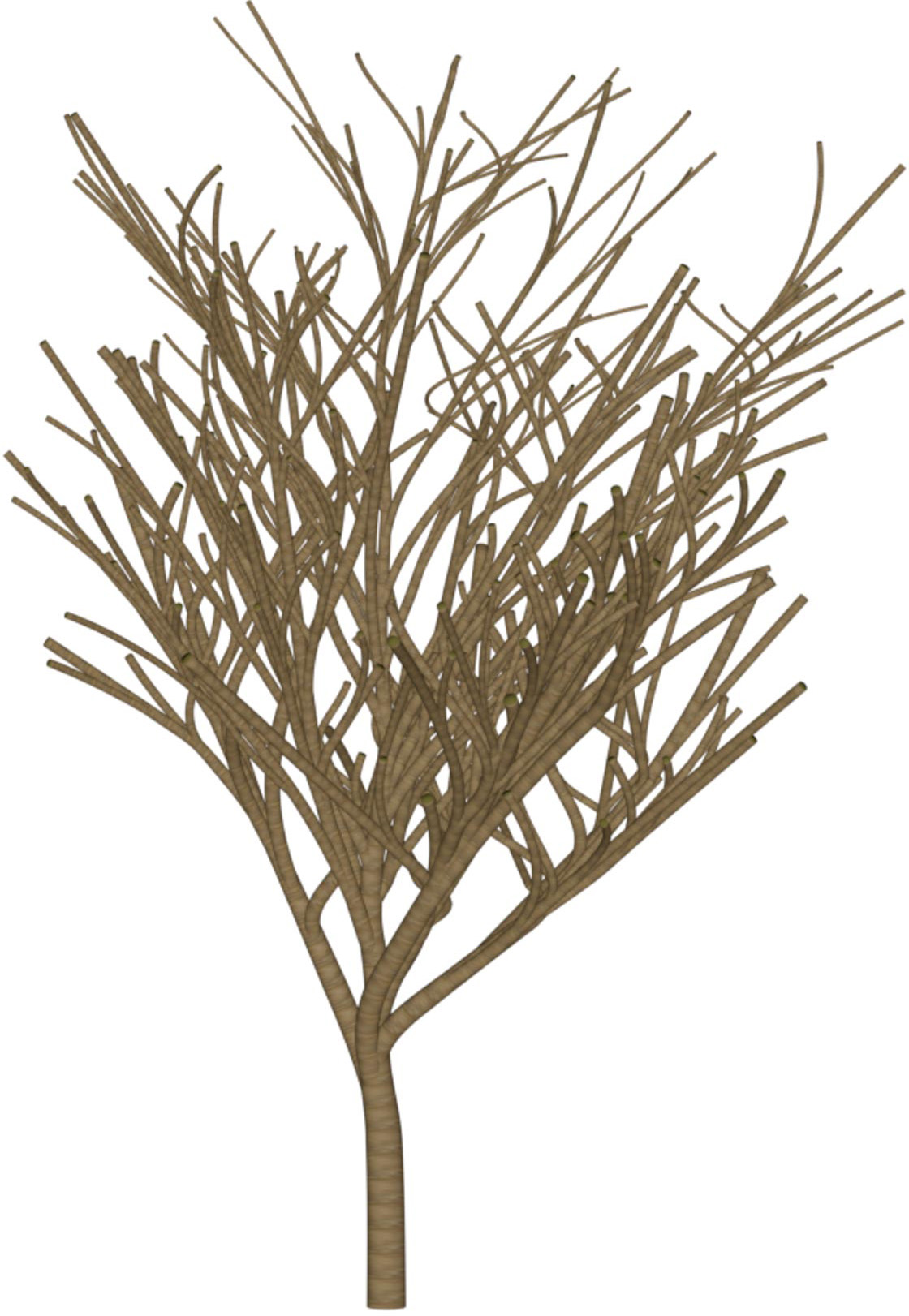
(a) Visual outputs for test trees.
| Method | NCD↓ | F1↑ | COV↑ |
|---|---|---|---|
| Tree D-Fusion | 0.66 | 0.01 | 16% |
| SVDTree | 0.31 | 0.58 | 72% |
| Latent L-Systems | 0.33 | 0.57 | 44% |
| TreeON | 0.24 | 0.76 | 86% |
| Ours | 0.21 | 0.81 | 75% |
(b) Quantitative results.
Table 2: Comparison with state-of-the-art methods.
| Inputs | Predictions | ||||
|---|---|---|---|---|---|
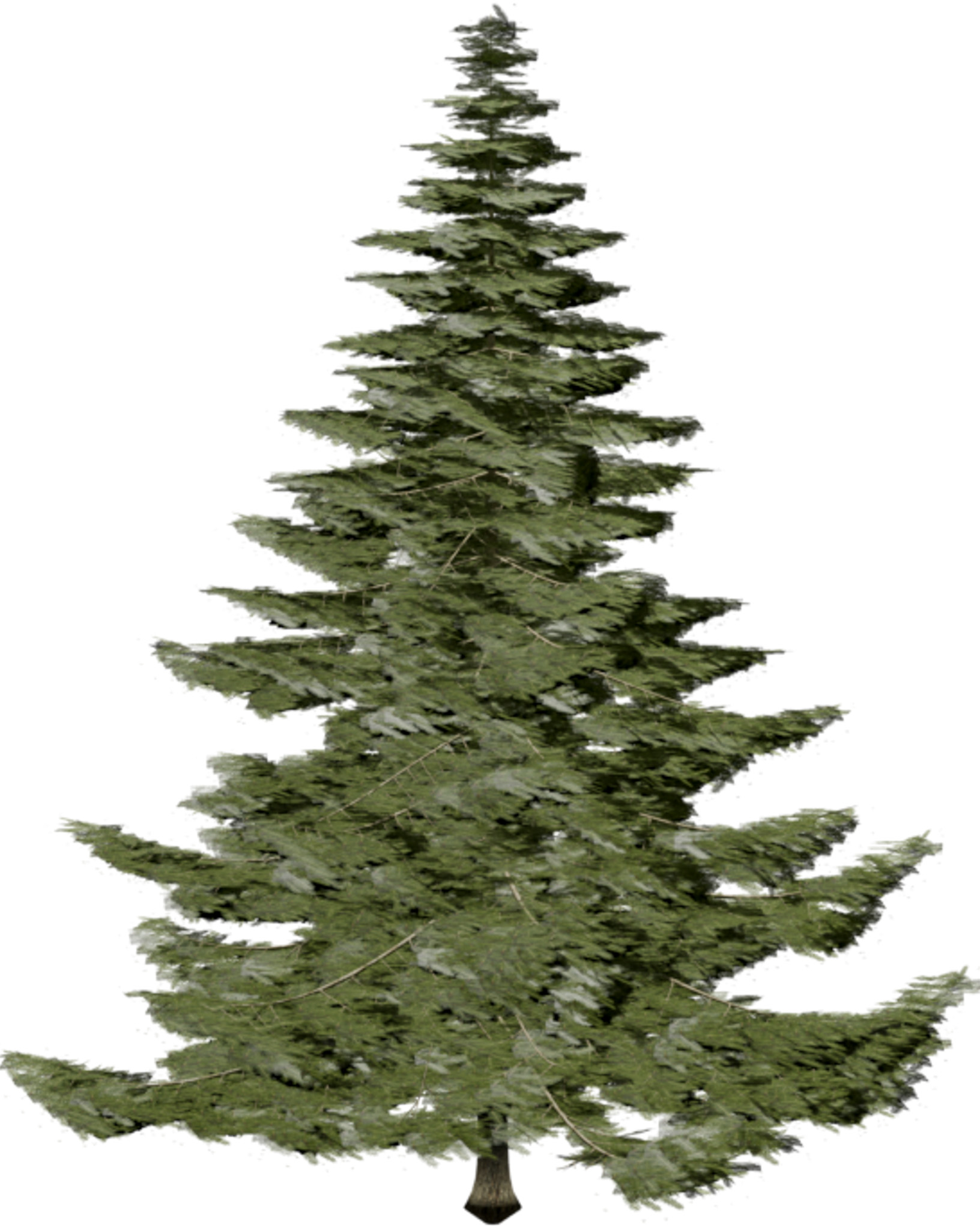 Target
Target
|
 DSM
DSM
|
 Orthophoto
Orthophoto
|
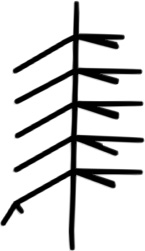 DSM only
DSM only
|
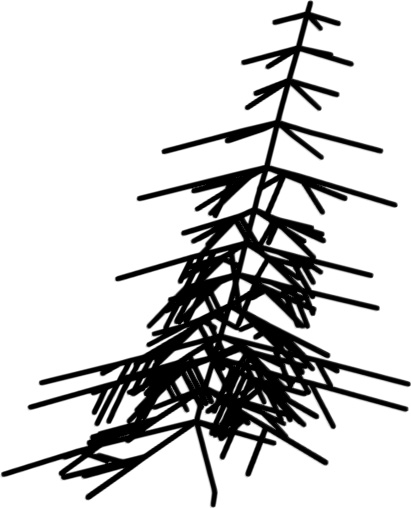 Ortho only
Ortho only
|
 DSM + Ortho
DSM + Ortho
|
Figure 2: Qualitative modality ablation on a representative tree.
| DSM | Orthophoto | LiDAR | Ours | |
|---|---|---|---|---|
| 1 |  |
 |
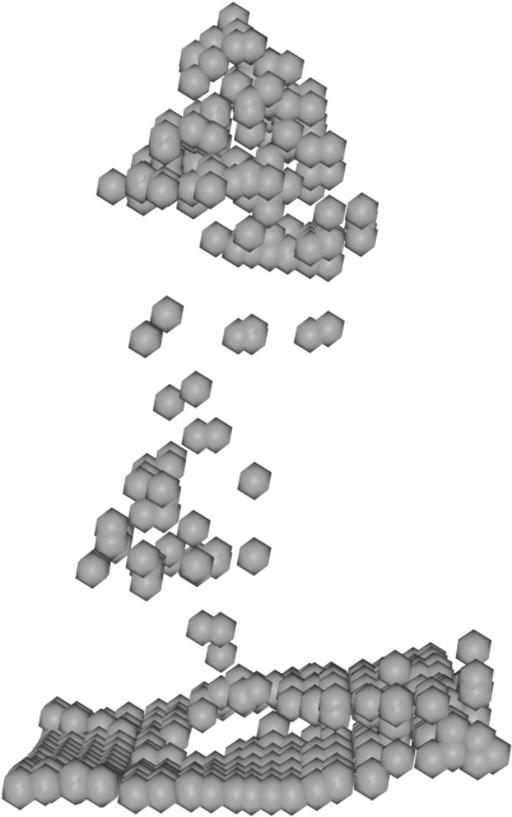 |
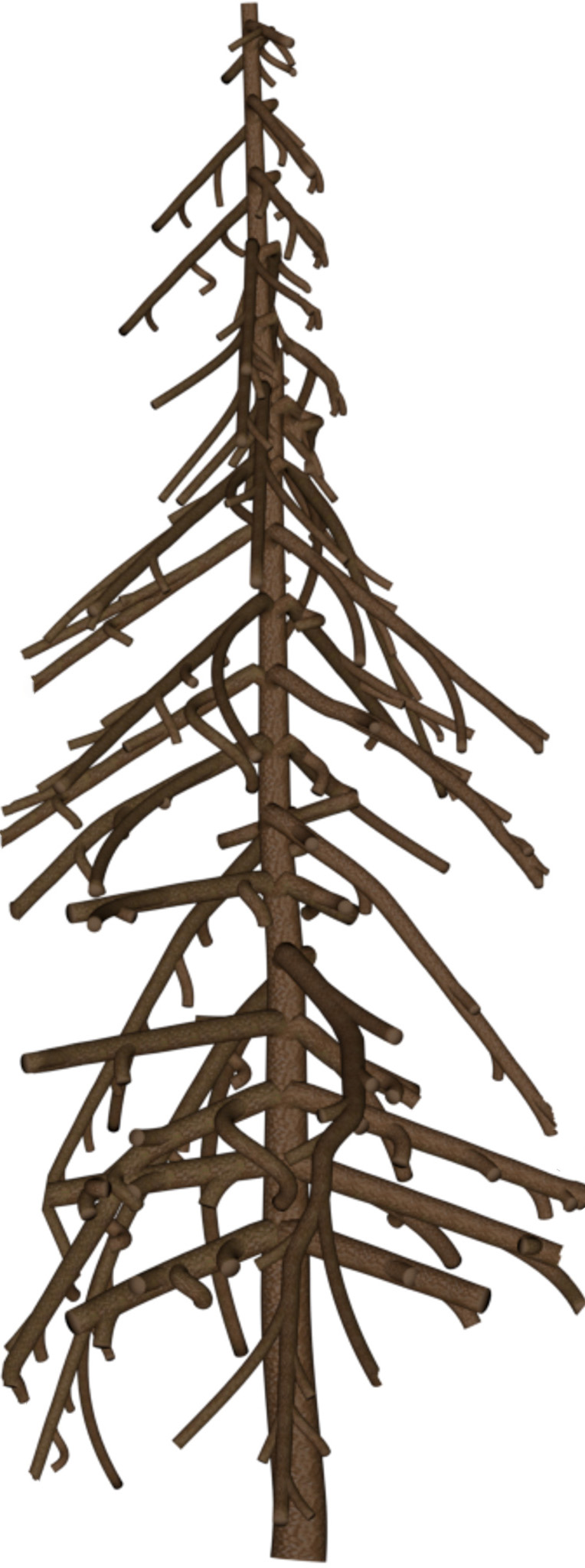 |
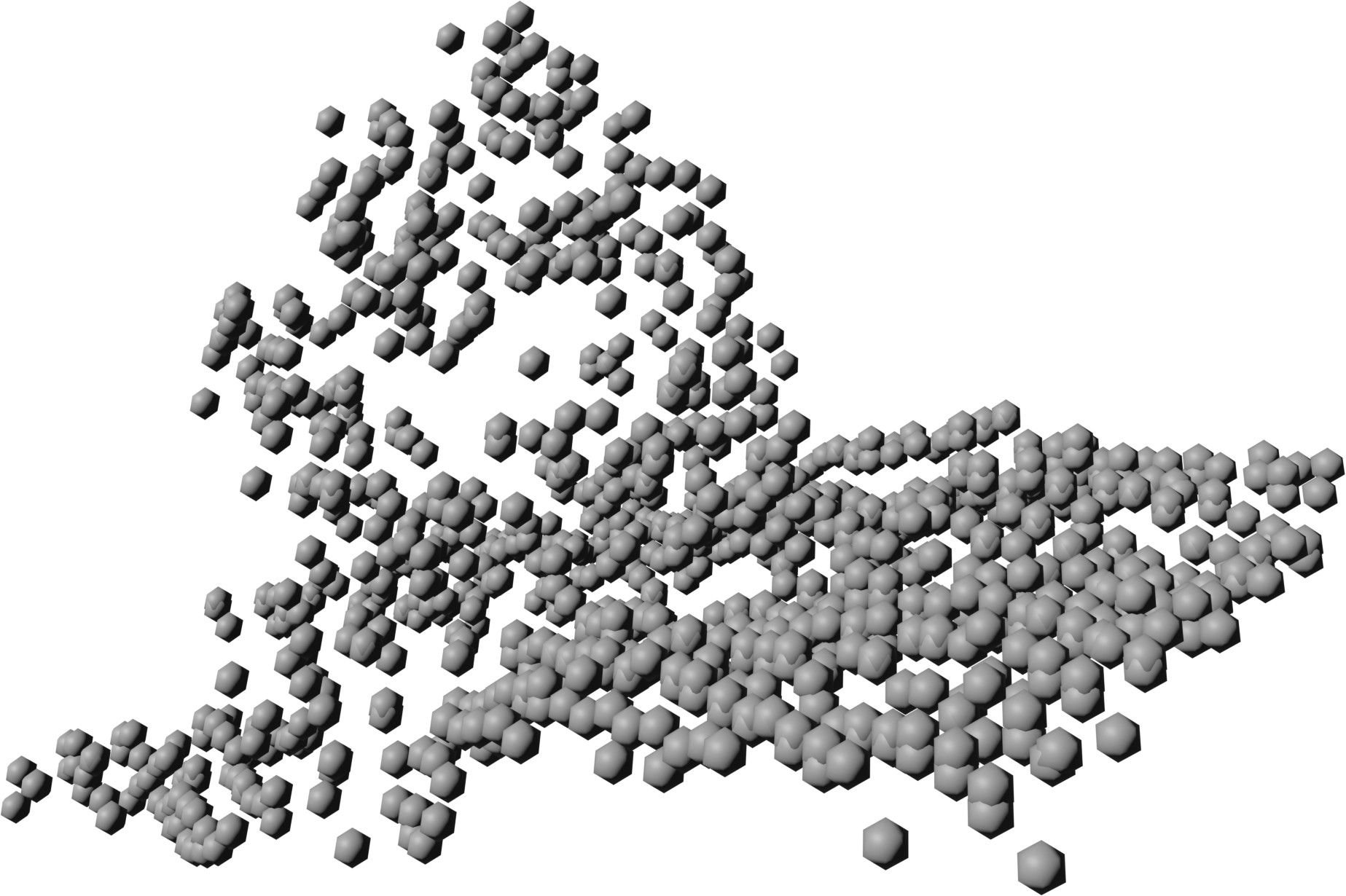
| 2 |  |
 |
 |
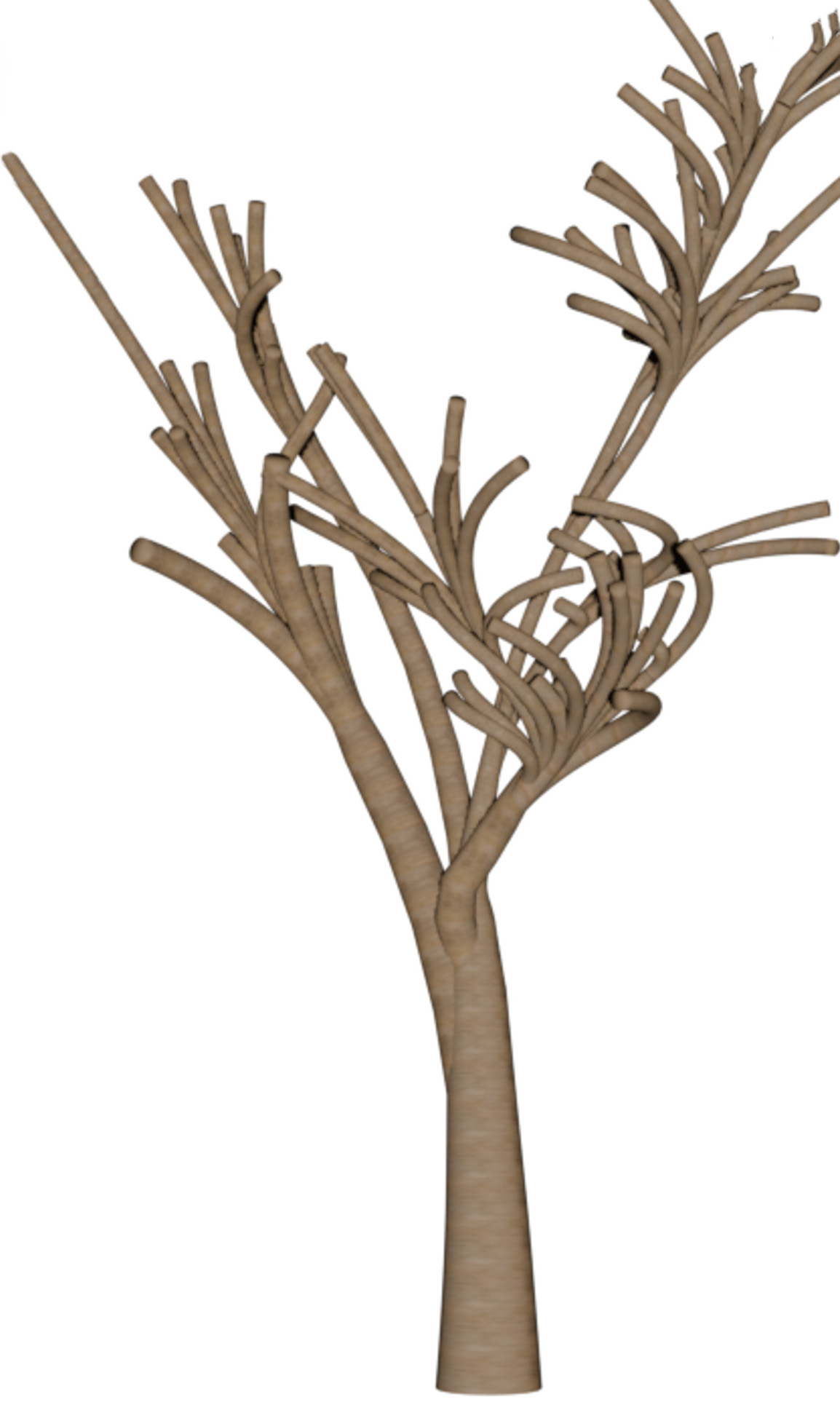 |
Table 3: Qualitative comparison against IGN LiDAR point clouds (37 and 25 pts/m²) in the French Pyrenees.